Expo
view channel
view channel
view channel
view channel
view channel
view channel
Medical Imaging
AI
Surgical TechniquesPatient CareHealth ITPoint of CareBusiness
Events
Webinars

- Rapid Clotting Gel Improves Emergency Bleeding Control
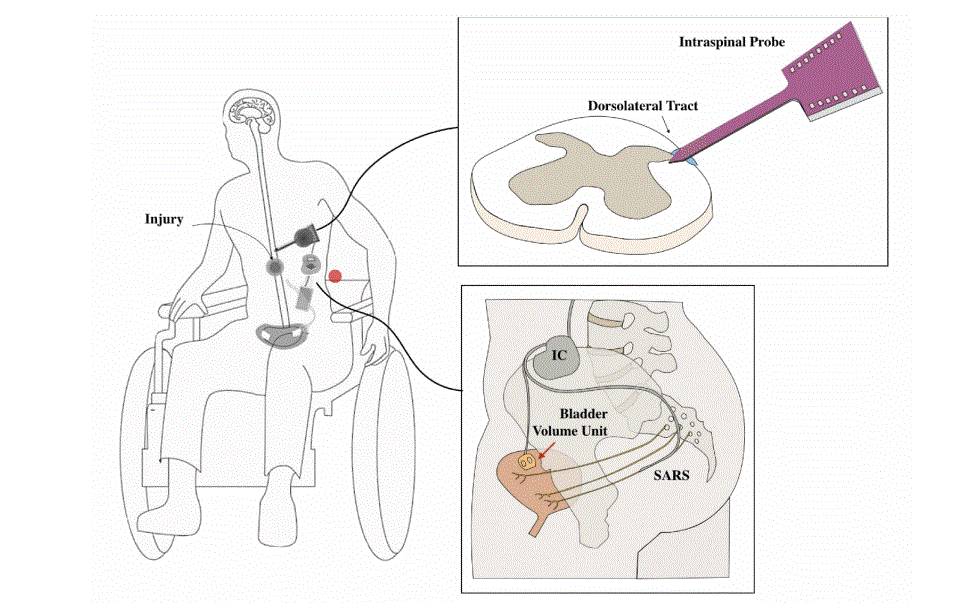
- Spinal Cord Interface Restores Bladder Control in Preclinical Study
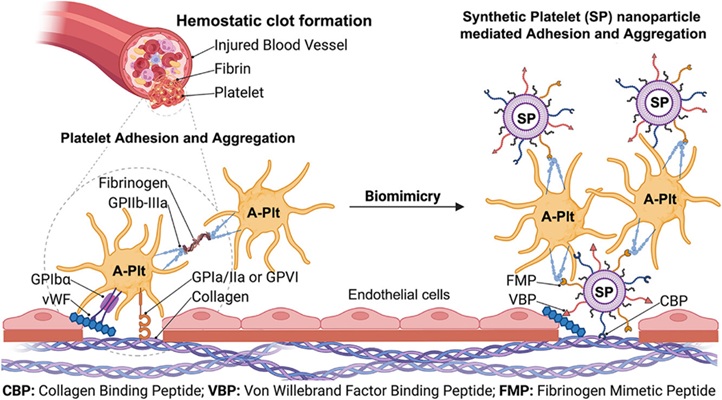
- Shelf-Stable Synthetic Platelets Could Enable Prehospital Hemorrhage Control
- Blood-Filtering Device May Prolong Pregnancy in Severe Early Preeclampsia
- AI Tool Predicts In-Hospital Cardiac Arrest Minutes in Advance
- Optical Wireless Link Enables Cable-Free 4K Endoscopic Imaging
- Glasses-Free 3D Imaging Software Receives FDA Clearance for Surgical Planning
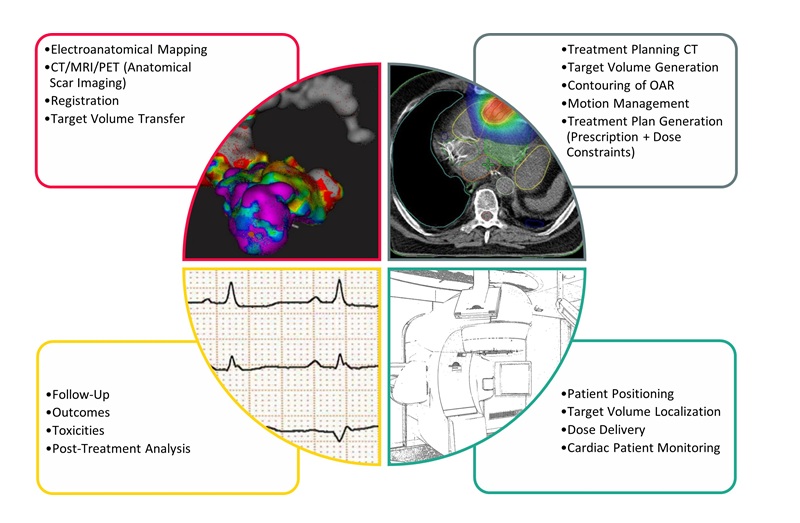
- Noninvasive Cardiac Radiotherapy Reduces Ventricular Tachycardia Events
- Augmented Reality System for Knee Replacement Receives FDA Clearance
- New AI Tool Predicts Complications Before Lung Cancer Surgery
- Revolutionary Automatic IV-Line Flushing Device to Enhance Infusion Care
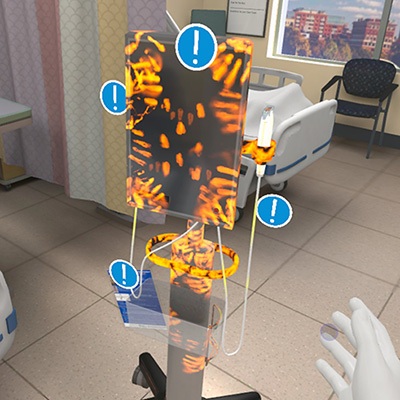
- VR Training Tool Combats Contamination of Portable Medical Equipment
- Portable Biosensor Platform to Reduce Hospital-Acquired Infections
- First-Of-Its-Kind Portable Germicidal Light Technology Disinfects High-Touch Clinical Surfaces in Seconds
- Surgical Capacity Optimization Solution Helps Hospitals Boost OR Utilization
- Proximie Advances AI-Driven Intelligent Operating Rooms with NVIDIA Collaboration
- GE HealthCare, DeepHealth Expand AI Breast Imaging Collaboration
- Sinocare Presents AI-Driven Integrated Digital Health Solutions at CMEF
- New Partnership Advances Physical AI into Perioperative Workflows
- GE HealthCare and Medtronic Expand Alliance with Intraoperative Imaging Integration
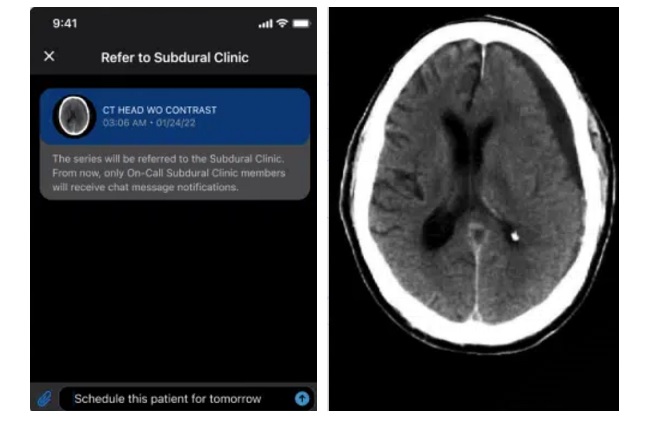
- AI System Detects and Quantifies Chronic Subdural Hematoma
- Continuous Monitoring Platform Detects Infection Risk Across Care Transitions
- Automated System Classifies and Tracks Cardiogenic Shock Across Hospital Settings
- Voice-Driven AI System Enables Structured GI Procedure Documentation
- EMR-Based Tool Predicts Graft Failure After Kidney Transplant
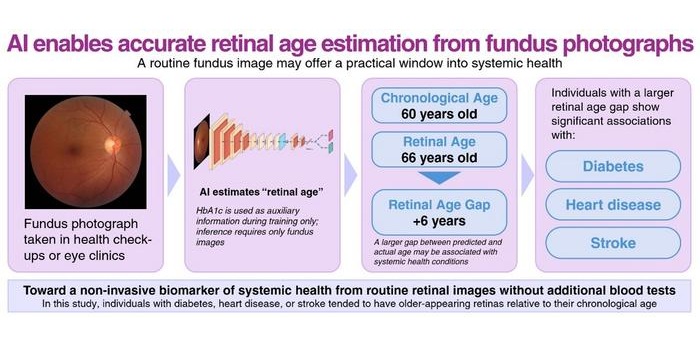
- AI Model Uses Eye Imaging to Identify Risk of Major Systemic Diseases
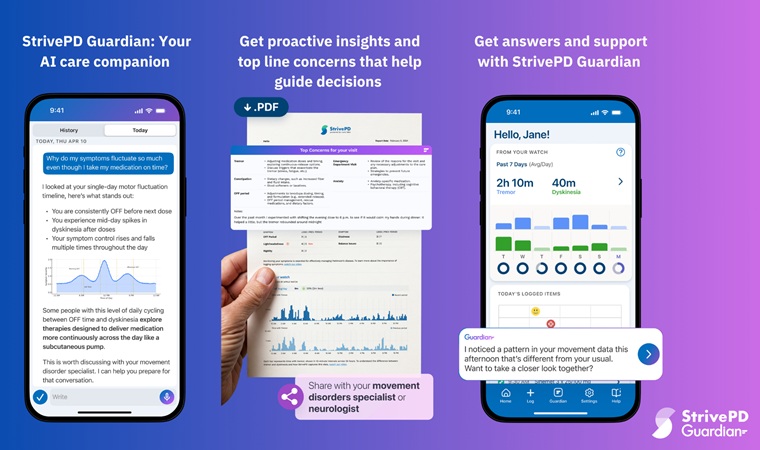
- AI Platform Interprets Real-Time Wearable Data for Parkinson’s Management
- Algorithm Identifies Cardiac Arrest Hotspots to Guide AED Placement
- AI Analysis of Pericardial Fat Refines Long-Term Heart Disease Risk
- Machine Learning Approach Enhances Liver Cancer Risk Stratification

 Expo
Expo
- Rapid Clotting Gel Improves Emergency Bleeding Control
- Spinal Cord Interface Restores Bladder Control in Preclinical Study
- Shelf-Stable Synthetic Platelets Could Enable Prehospital Hemorrhage Control
- Blood-Filtering Device May Prolong Pregnancy in Severe Early Preeclampsia
- AI Tool Predicts In-Hospital Cardiac Arrest Minutes in Advance
- Optical Wireless Link Enables Cable-Free 4K Endoscopic Imaging
- Glasses-Free 3D Imaging Software Receives FDA Clearance for Surgical Planning
- Noninvasive Cardiac Radiotherapy Reduces Ventricular Tachycardia Events
- Augmented Reality System for Knee Replacement Receives FDA Clearance
- New AI Tool Predicts Complications Before Lung Cancer Surgery
- Revolutionary Automatic IV-Line Flushing Device to Enhance Infusion Care
- VR Training Tool Combats Contamination of Portable Medical Equipment
- Portable Biosensor Platform to Reduce Hospital-Acquired Infections
- First-Of-Its-Kind Portable Germicidal Light Technology Disinfects High-Touch Clinical Surfaces in Seconds
- Surgical Capacity Optimization Solution Helps Hospitals Boost OR Utilization
- Proximie Advances AI-Driven Intelligent Operating Rooms with NVIDIA Collaboration
- GE HealthCare, DeepHealth Expand AI Breast Imaging Collaboration
- Sinocare Presents AI-Driven Integrated Digital Health Solutions at CMEF
- New Partnership Advances Physical AI into Perioperative Workflows
- GE HealthCare and Medtronic Expand Alliance with Intraoperative Imaging Integration
- AI System Detects and Quantifies Chronic Subdural Hematoma
- Continuous Monitoring Platform Detects Infection Risk Across Care Transitions
- Automated System Classifies and Tracks Cardiogenic Shock Across Hospital Settings
- Voice-Driven AI System Enables Structured GI Procedure Documentation
- EMR-Based Tool Predicts Graft Failure After Kidney Transplant
- AI Model Uses Eye Imaging to Identify Risk of Major Systemic Diseases
- AI Platform Interprets Real-Time Wearable Data for Parkinson’s Management
- Algorithm Identifies Cardiac Arrest Hotspots to Guide AED Placement
- AI Analysis of Pericardial Fat Refines Long-Term Heart Disease Risk
- Machine Learning Approach Enhances Liver Cancer Risk Stratification