Expo
view channel
view channel
view channel
view channel
view channel
view channel
Medical Imaging
AICritical CareSurgical TechniquesPatient CareHealth ITPoint of CareBusiness
Events
Webinars

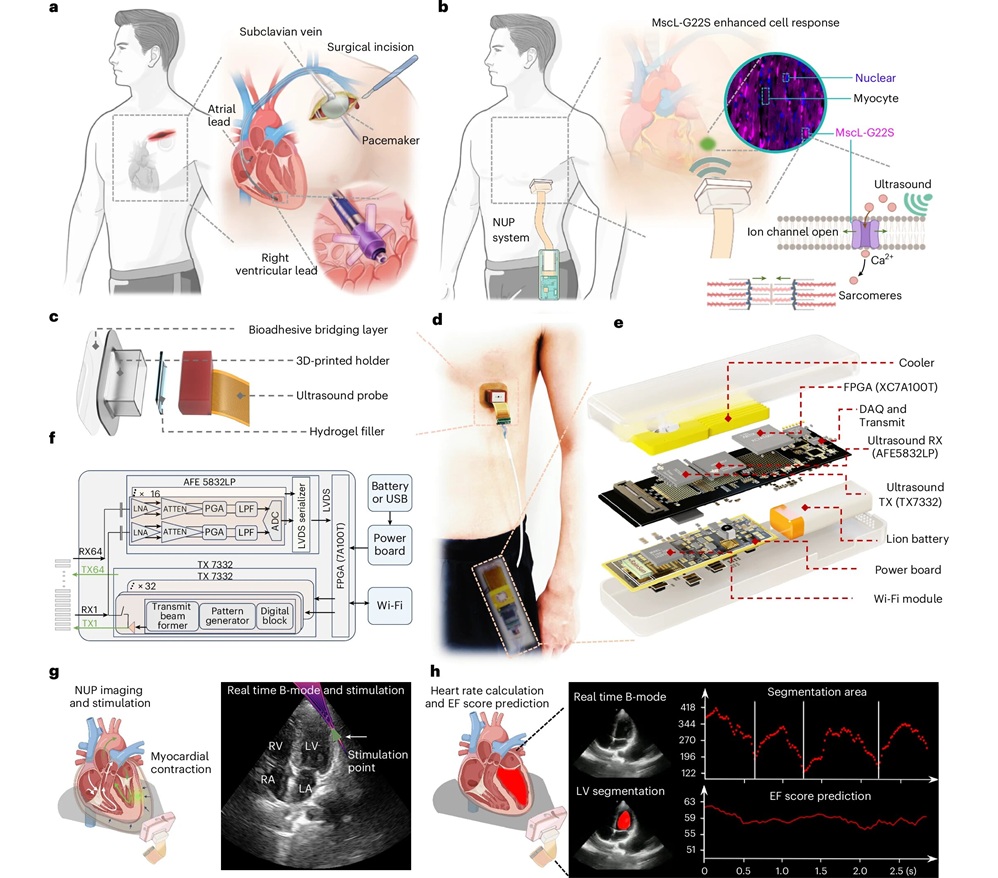
- Wearable Ultrasound Patch Noninvasively Paces Heart to Stabilize Arrhythmias
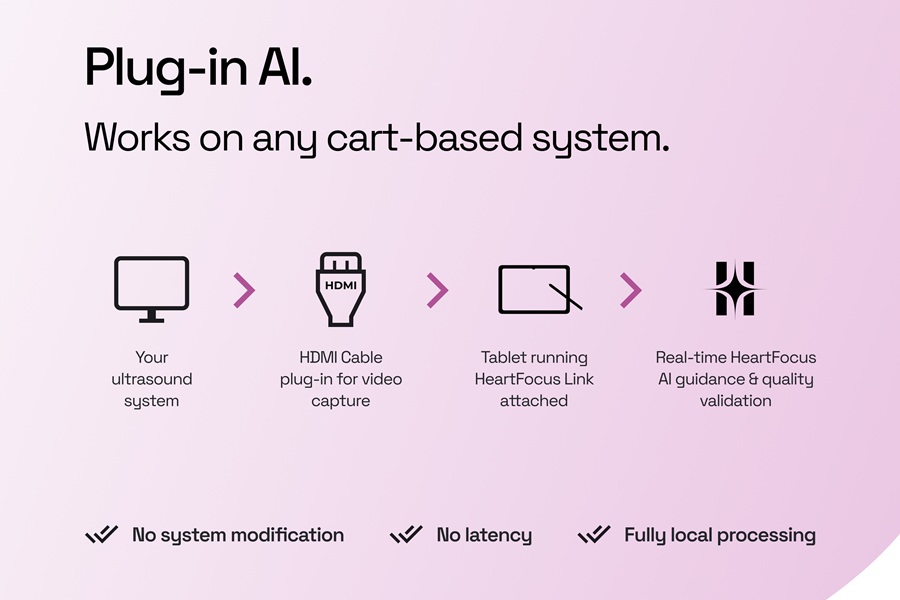
- AI-Guided System Supports Cardiac Ultrasound Training on Cart-Based Systems
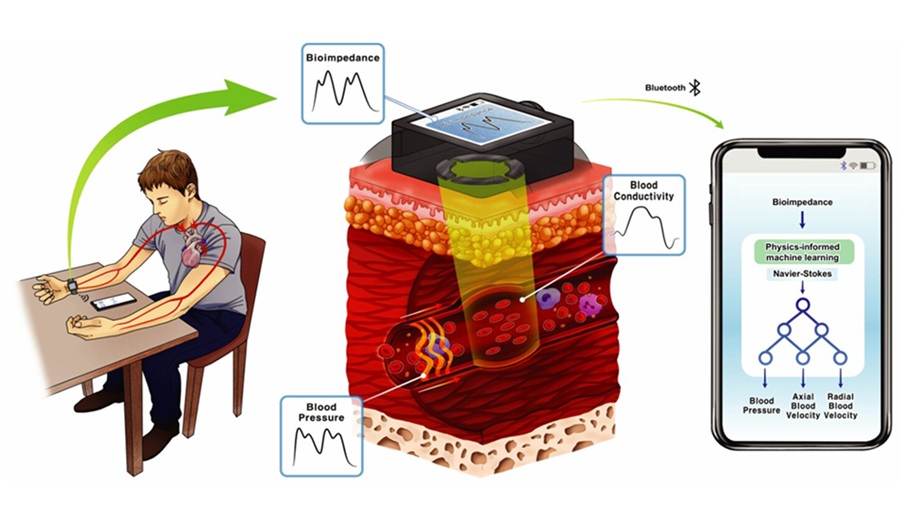
- Cuffless Wearable Enables Continuous Blood Pressure Monitoring for Hypertension Care
- AI ECG Tool Detects Cardiac Amyloidosis for Early Screening
- AI ECG Index Tracks Pubertal Maturation in Children and Adolescents
- Small Cryoprobe Outperforms Forceps in Lung Biopsy Trial
- Natural Bypass Score May Guide Care in Chronic Coronary Blockages
- Navigation Instruments Cleared for Posterior Cervical Fusion Procedures
- Bioengineered Heart Patch Improves Cardiac Function in Advanced Heart Failure
- Fracture Plating System Combines Anatomical Fit with Streamlined Instrumentation
- Wearable Sleep Data Predict Adherence to Pulmonary Rehabilitation
- Revolutionary Automatic IV-Line Flushing Device to Enhance Infusion Care
- VR Training Tool Combats Contamination of Portable Medical Equipment
- Portable Biosensor Platform to Reduce Hospital-Acquired Infections
- First-Of-Its-Kind Portable Germicidal Light Technology Disinfects High-Touch Clinical Surfaces in Seconds
- Artivion Adds FDA-Approved NEXUS System to Aortic Arch Portfolio
- Olympus Partnership Aims to Expand Access to Robot-Assisted Endoscopic Therapy
- Johnson & Johnson Launches AI-Driven Cardiac Mapping System
- Proximie Advances AI-Driven Intelligent Operating Rooms with NVIDIA Collaboration
- GE HealthCare, DeepHealth Expand AI Breast Imaging Collaboration
- EHR-Integrated Screening Workflow Detects Cognitive Impairment at Admission
- AI System Detects and Quantifies Chronic Subdural Hematoma
- Continuous Monitoring Platform Detects Infection Risk Across Care Transitions
- Automated System Classifies and Tracks Cardiogenic Shock Across Hospital Settings
- Voice-Driven AI System Enables Structured GI Procedure Documentation
- AI Tool Predicts Unplanned Care and Symptom Burden in Cancer Survivors
- Automated Phone Speech Test Identifies Alzheimer’s Pathology for Prescreening
- FDA-Cleared AI System Detects Sepsis Earlier and Reduces Mortality
- Facial Image Analysis Tracks Biological Aging, Predicts Cancer Outcomes
- AI Model Uses Eye Imaging to Identify Risk of Major Systemic Diseases

 Expo
Expo
- Wearable Ultrasound Patch Noninvasively Paces Heart to Stabilize Arrhythmias
- AI-Guided System Supports Cardiac Ultrasound Training on Cart-Based Systems
- Cuffless Wearable Enables Continuous Blood Pressure Monitoring for Hypertension Care
- AI ECG Tool Detects Cardiac Amyloidosis for Early Screening
- AI ECG Index Tracks Pubertal Maturation in Children and Adolescents
- Small Cryoprobe Outperforms Forceps in Lung Biopsy Trial
- Natural Bypass Score May Guide Care in Chronic Coronary Blockages
- Navigation Instruments Cleared for Posterior Cervical Fusion Procedures
- Bioengineered Heart Patch Improves Cardiac Function in Advanced Heart Failure
- Fracture Plating System Combines Anatomical Fit with Streamlined Instrumentation
- Wearable Sleep Data Predict Adherence to Pulmonary Rehabilitation
- Revolutionary Automatic IV-Line Flushing Device to Enhance Infusion Care
- VR Training Tool Combats Contamination of Portable Medical Equipment
- Portable Biosensor Platform to Reduce Hospital-Acquired Infections
- First-Of-Its-Kind Portable Germicidal Light Technology Disinfects High-Touch Clinical Surfaces in Seconds
- Artivion Adds FDA-Approved NEXUS System to Aortic Arch Portfolio
- Olympus Partnership Aims to Expand Access to Robot-Assisted Endoscopic Therapy
- Johnson & Johnson Launches AI-Driven Cardiac Mapping System
- Proximie Advances AI-Driven Intelligent Operating Rooms with NVIDIA Collaboration
- GE HealthCare, DeepHealth Expand AI Breast Imaging Collaboration
- EHR-Integrated Screening Workflow Detects Cognitive Impairment at Admission
- AI System Detects and Quantifies Chronic Subdural Hematoma
- Continuous Monitoring Platform Detects Infection Risk Across Care Transitions
- Automated System Classifies and Tracks Cardiogenic Shock Across Hospital Settings
- Voice-Driven AI System Enables Structured GI Procedure Documentation
- AI Tool Predicts Unplanned Care and Symptom Burden in Cancer Survivors
- Automated Phone Speech Test Identifies Alzheimer’s Pathology for Prescreening
- FDA-Cleared AI System Detects Sepsis Earlier and Reduces Mortality
- Facial Image Analysis Tracks Biological Aging, Predicts Cancer Outcomes
- AI Model Uses Eye Imaging to Identify Risk of Major Systemic Diseases