Expo
view channel
view channel
view channel
view channel
view channel
view channel
Medical Imaging
AICritical CareSurgical TechniquesPatient CareHealth ITPoint of CareBusiness
Events
Webinars

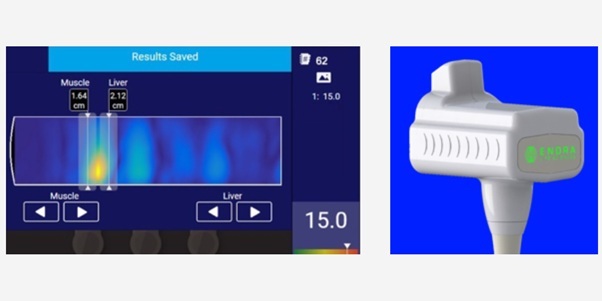
- Portable Ultrasound Tool Quantifies Liver Fat with MRI-Like Accuracy
- AI Method Turns Toe Scan into Rapid PAD Screening Tool
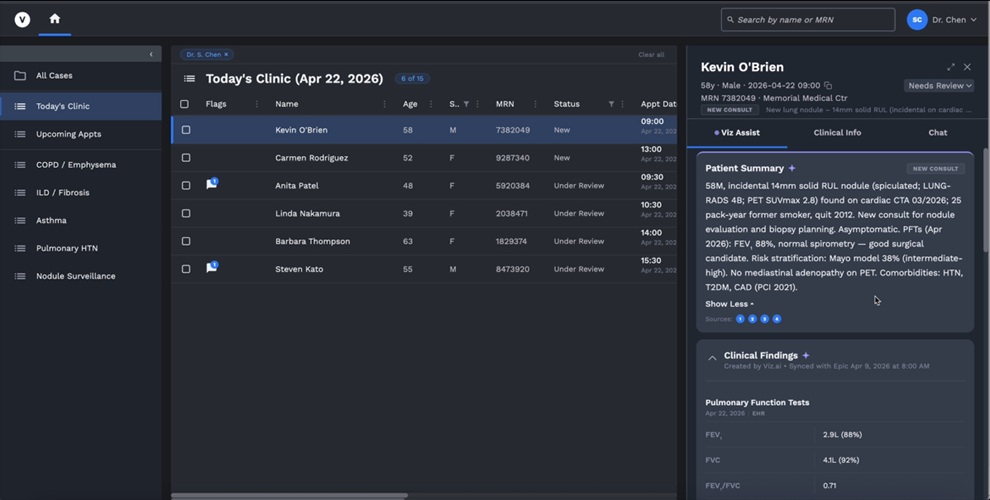
- Integrated AI Pulmonary Workflow System Streamlines Detection and Follow-Up
- AI Model Predicts 10-Year Stroke Risk from Standard ECG
- Implantable Cytokine Device Enables Localized Immunotherapy for Ovarian Cancer
- FDA-Cleared Nerve Stimulator Advances Intraoperative Peripheral Nerve Assessment
- Handheld AI Endomicroscope Enables Real-Time Precancer Detection at Point of Care
- Intravascular Lithotripsy Catheter Advances Treatment of Calcified Coronary Disease
- Advanced Endoscopy Platform Targets Challenging Upper GI Procedures
- Photoacoustic Imaging System Maps Hidden Nerves and Vessels in Robotic Laparoscopy
- Wearable Sleep Data Predict Adherence to Pulmonary Rehabilitation
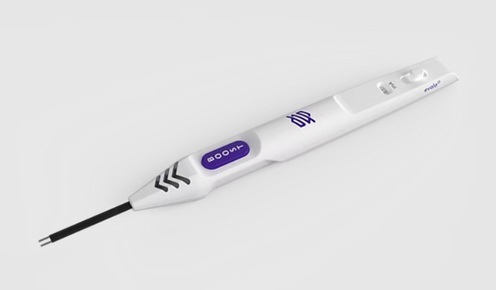
- Revolutionary Automatic IV-Line Flushing Device to Enhance Infusion Care
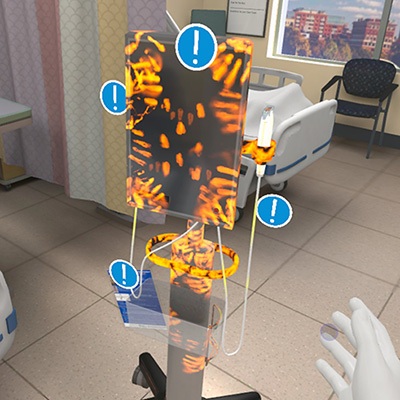
- VR Training Tool Combats Contamination of Portable Medical Equipment
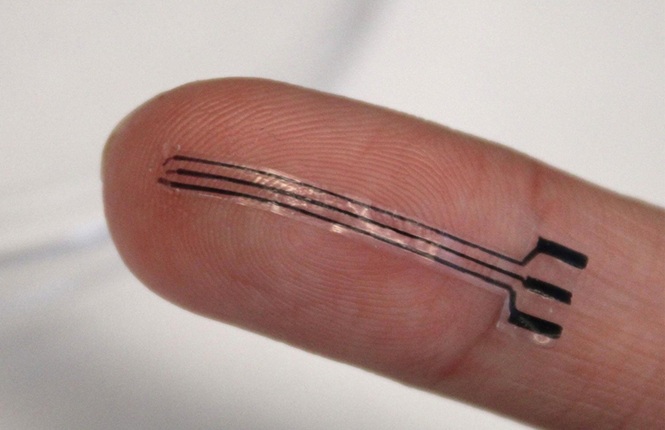
- Portable Biosensor Platform to Reduce Hospital-Acquired Infections
- First-Of-Its-Kind Portable Germicidal Light Technology Disinfects High-Touch Clinical Surfaces in Seconds
- Johnson & Johnson Launches AI-Driven Cardiac Mapping System
- Proximie Advances AI-Driven Intelligent Operating Rooms with NVIDIA Collaboration
- GE HealthCare, DeepHealth Expand AI Breast Imaging Collaboration
- Sinocare Presents AI-Driven Integrated Digital Health Solutions at CMEF
- New Partnership Advances Physical AI into Perioperative Workflows
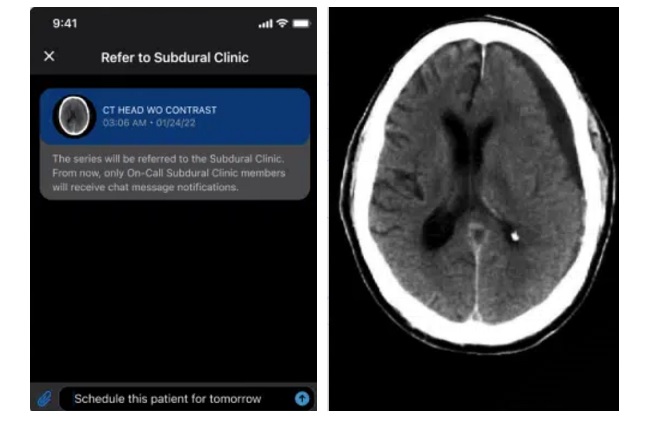
- AI System Detects and Quantifies Chronic Subdural Hematoma
- Continuous Monitoring Platform Detects Infection Risk Across Care Transitions
- Automated System Classifies and Tracks Cardiogenic Shock Across Hospital Settings
- Voice-Driven AI System Enables Structured GI Procedure Documentation
- EMR-Based Tool Predicts Graft Failure After Kidney Transplant
- Facial Image Analysis Tracks Biological Aging, Predicts Cancer Outcomes
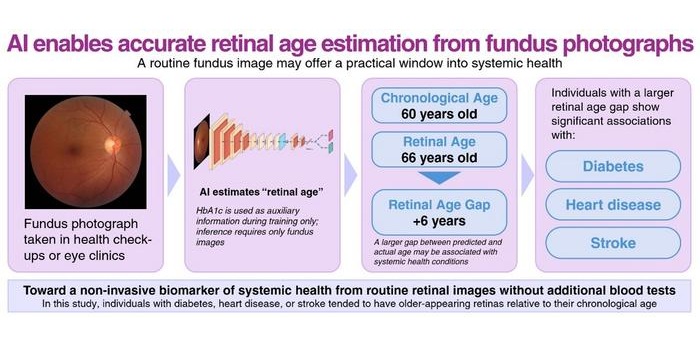
- AI Model Uses Eye Imaging to Identify Risk of Major Systemic Diseases
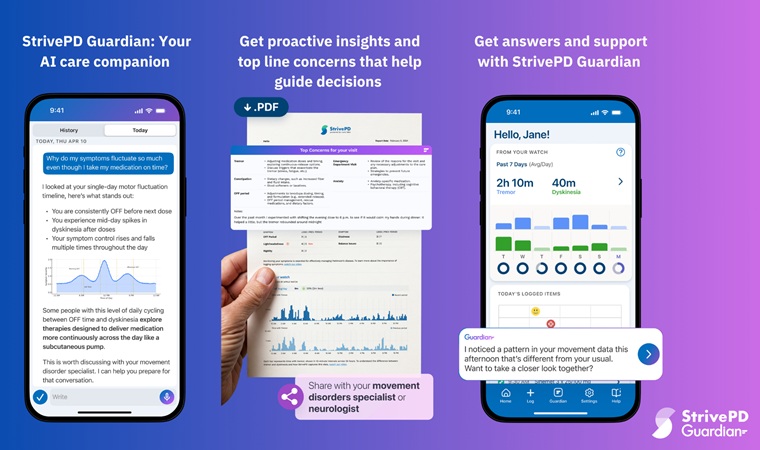
- AI Platform Interprets Real-Time Wearable Data for Parkinson’s Management
- Algorithm Identifies Cardiac Arrest Hotspots to Guide AED Placement
- AI Analysis of Pericardial Fat Refines Long-Term Heart Disease Risk

 Expo
Expo
- Portable Ultrasound Tool Quantifies Liver Fat with MRI-Like Accuracy
- AI Method Turns Toe Scan into Rapid PAD Screening Tool
- Integrated AI Pulmonary Workflow System Streamlines Detection and Follow-Up
- AI Model Predicts 10-Year Stroke Risk from Standard ECG
- Implantable Cytokine Device Enables Localized Immunotherapy for Ovarian Cancer
- FDA-Cleared Nerve Stimulator Advances Intraoperative Peripheral Nerve Assessment
- Handheld AI Endomicroscope Enables Real-Time Precancer Detection at Point of Care
- Intravascular Lithotripsy Catheter Advances Treatment of Calcified Coronary Disease
- Advanced Endoscopy Platform Targets Challenging Upper GI Procedures
- Photoacoustic Imaging System Maps Hidden Nerves and Vessels in Robotic Laparoscopy
- Wearable Sleep Data Predict Adherence to Pulmonary Rehabilitation
- Revolutionary Automatic IV-Line Flushing Device to Enhance Infusion Care
- VR Training Tool Combats Contamination of Portable Medical Equipment
- Portable Biosensor Platform to Reduce Hospital-Acquired Infections
- First-Of-Its-Kind Portable Germicidal Light Technology Disinfects High-Touch Clinical Surfaces in Seconds
- Johnson & Johnson Launches AI-Driven Cardiac Mapping System
- Proximie Advances AI-Driven Intelligent Operating Rooms with NVIDIA Collaboration
- GE HealthCare, DeepHealth Expand AI Breast Imaging Collaboration
- Sinocare Presents AI-Driven Integrated Digital Health Solutions at CMEF
- New Partnership Advances Physical AI into Perioperative Workflows
- AI System Detects and Quantifies Chronic Subdural Hematoma
- Continuous Monitoring Platform Detects Infection Risk Across Care Transitions
- Automated System Classifies and Tracks Cardiogenic Shock Across Hospital Settings
- Voice-Driven AI System Enables Structured GI Procedure Documentation
- EMR-Based Tool Predicts Graft Failure After Kidney Transplant
- Facial Image Analysis Tracks Biological Aging, Predicts Cancer Outcomes
- AI Model Uses Eye Imaging to Identify Risk of Major Systemic Diseases
- AI Platform Interprets Real-Time Wearable Data for Parkinson’s Management
- Algorithm Identifies Cardiac Arrest Hotspots to Guide AED Placement
- AI Analysis of Pericardial Fat Refines Long-Term Heart Disease Risk